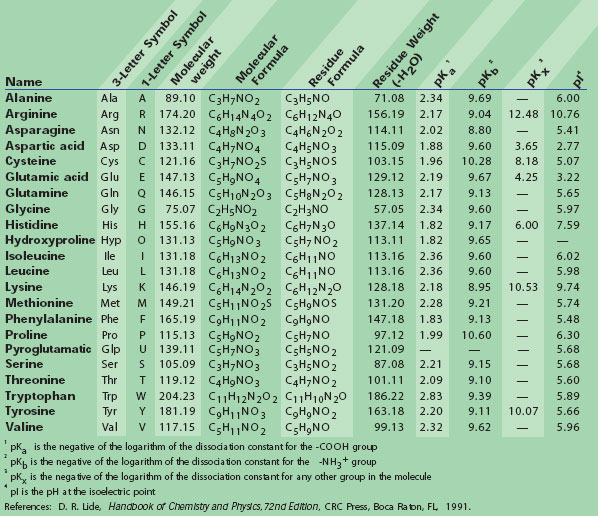
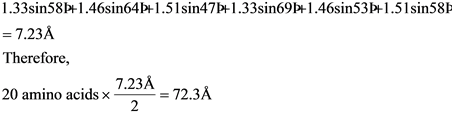Amyloid-like amelogenin nanoribbons template mineralization via a low-energy interface of ion binding sites | PNAS
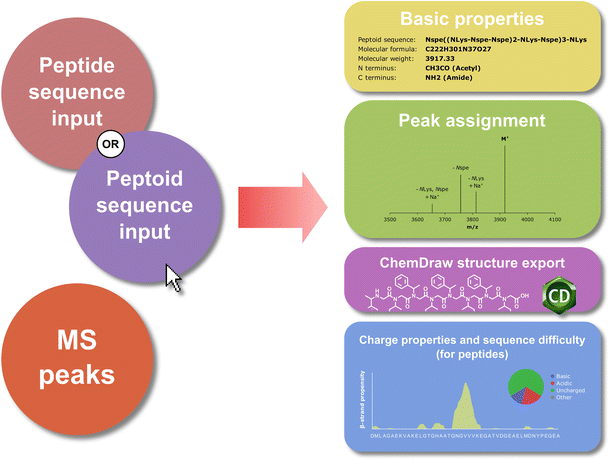
Pep-Calc.com: a set of web utilities for the calculation of peptide and peptoid properties and automatic mass spectral peak assignment | SpringerLink
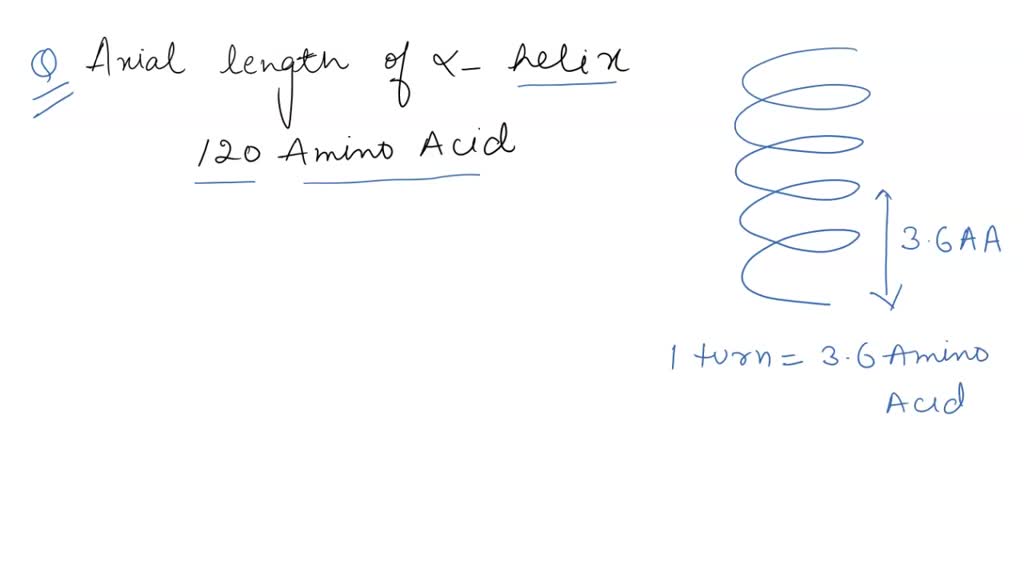
SOLVED: Calculate the axial length of an alpha helix that is of 120 amino acids long? How long would the polypeptide be if it were fully extended?

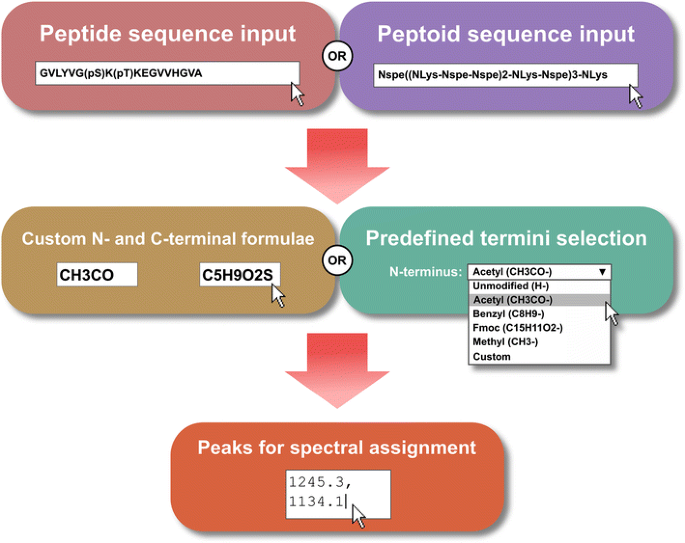
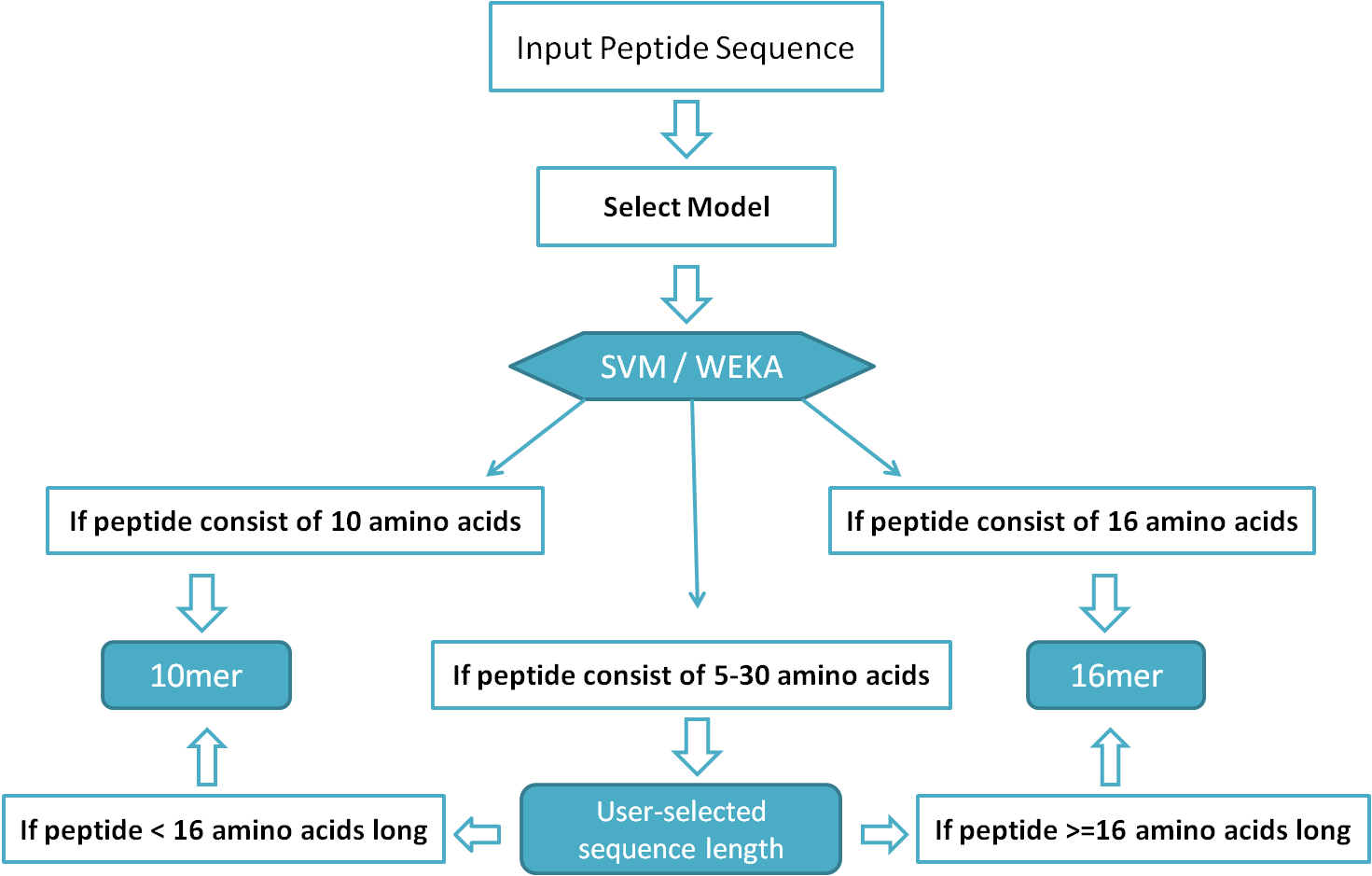
Peptide calculator inputs and outputs. a Summary of the input to the... | Download Scientific Diagram
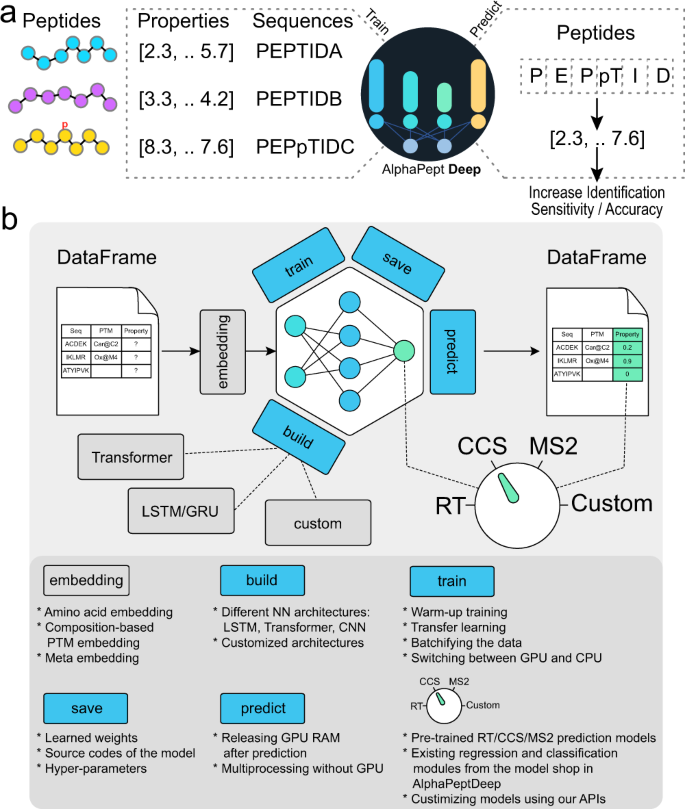
AlphaPeptDeep: a modular deep learning framework to predict peptide properties for proteomics | Nature Communications

Peptide calculator inputs and outputs. a Summary of the input to the... | Download Scientific Diagram

Targeted Quantification of Peptides Using Miniature Mass Spectrometry | Journal of Proteome Research

Solubility test of NSP and NSP r. (A) Peptide models computed by the... | Download Scientific Diagram

Sequence-Specific Model for Predicting Peptide Collision Cross Section Values in Proteomic Ion Mobility Spectrometry | Journal of Proteome Research

IJMS | Free Full-Text | Computational Evolution of Beta-2-Microglobulin Binding Peptides for Nanopatterned Surface Sensors