Screenshot of the homepage of the Huangjiu Yeast Genome Database and... | Download Scientific Diagram
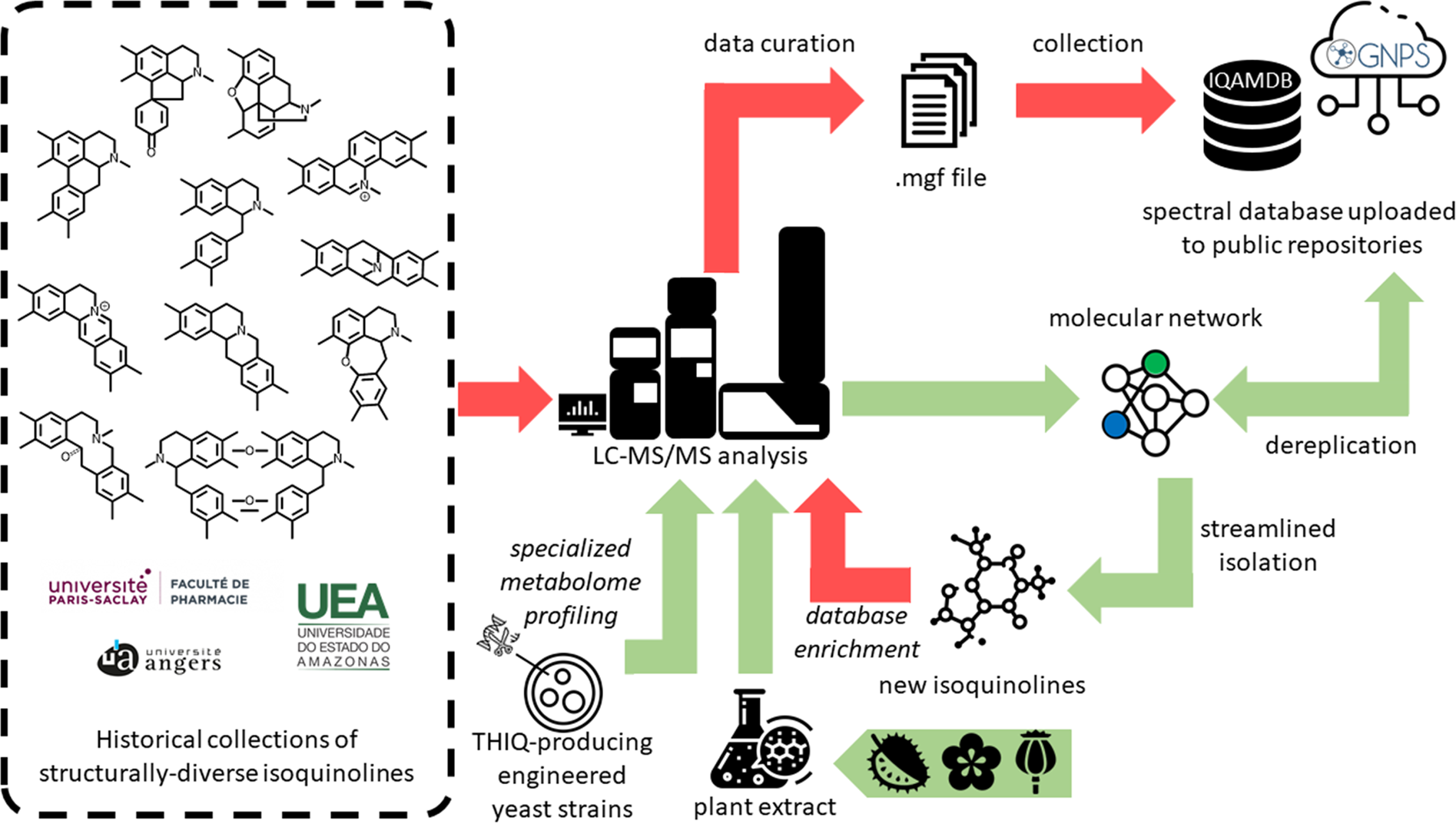
Implementation of a MS/MS database for isoquinoline alkaloids and other annonaceous metabolites | Scientific Data

Lallemand Inc DB-R43Y-0P5D Nottingham Ale Yeast(11 grams): Leaveners And Yeasts: Amazon.com: Industrial & Scientific

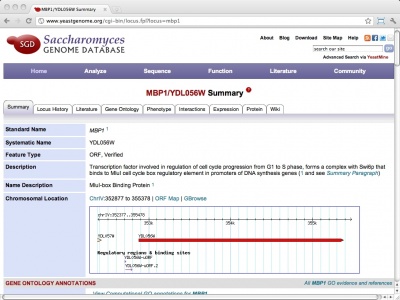
Yeast Protein database (YPD): a database for the complete proteome of Saccharomyces cerevisiae | Semantic Scholar

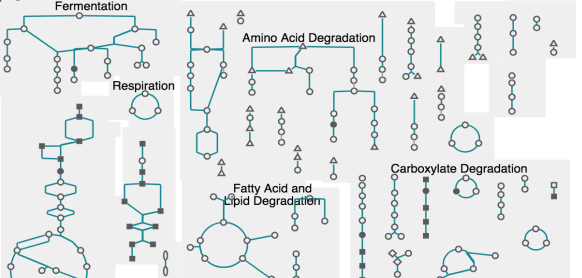
A consensus S. cerevisiae metabolic model Yeast8 and its ecosystem for comprehensively probing cellular metabolism | Nature Communications
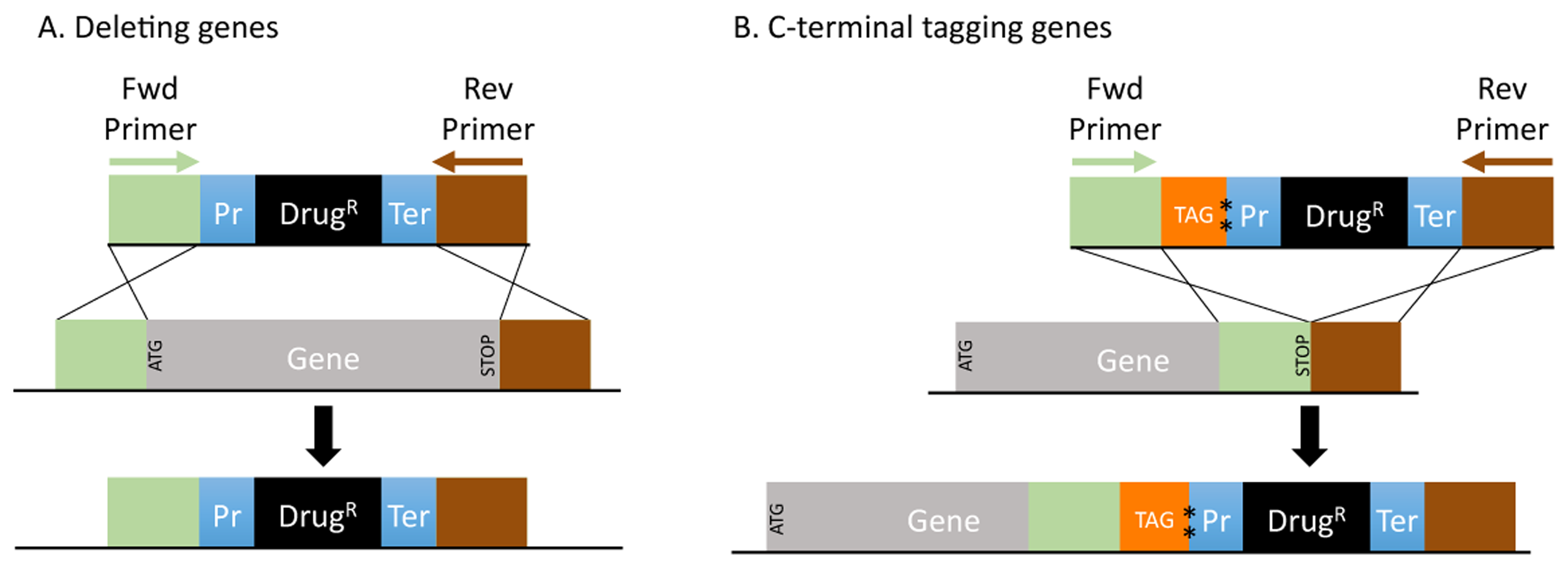
PRIMED: PRIMEr Database for Deleting and Tagging All Fission and Budding Yeast Genes Developed Using the Open-Source Genome Retrieval Script (GRS) - Open.ConductScience
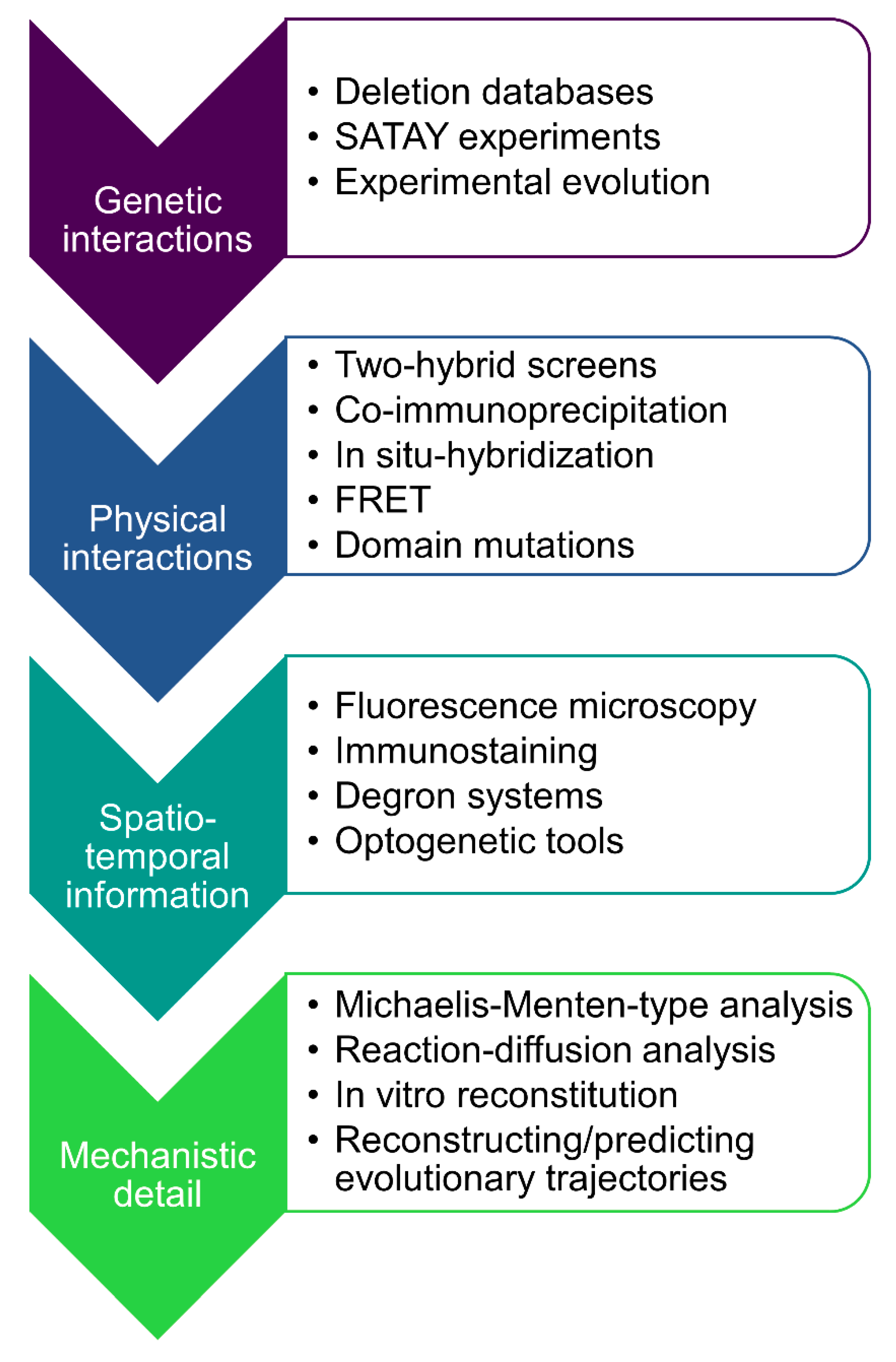
Cells | Free Full-Text | The Path towards Predicting Evolution as Illustrated in Yeast Cell Polarity
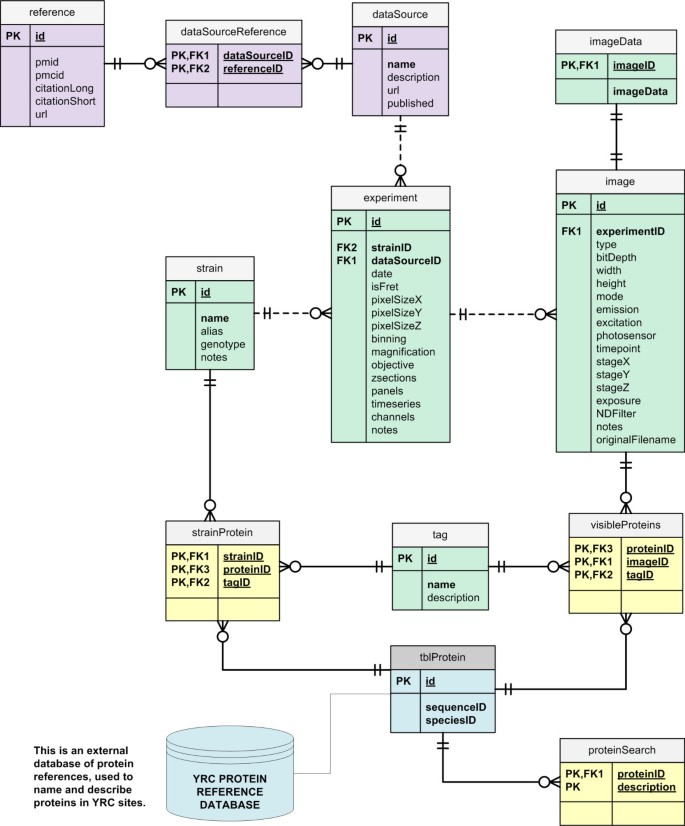
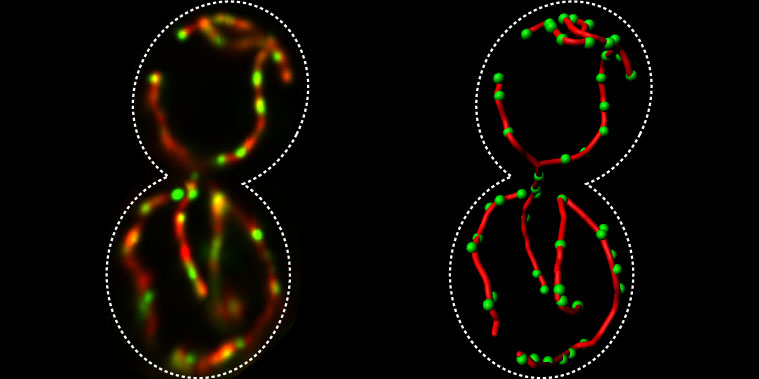
The Yeast Resource Center Public Image Repository: A large database of fluorescence microscopy images | BMC Bioinformatics | Full Text

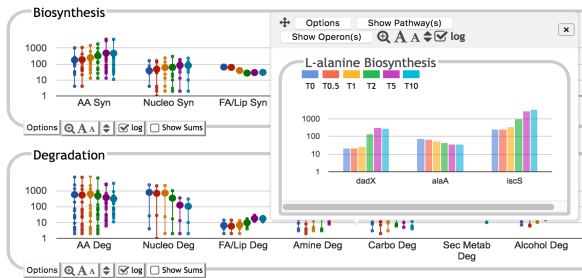
![PDF] YMDB: the Yeast Metabolome Database | Semantic Scholar PDF] YMDB: the Yeast Metabolome Database | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/b27c179caa2ec55fd3205f75e146713b5e073a3f/4-Figure1-1.png)